Biological Measurement Can Now Scale.
Biological measurement never achieved a Moore’s Law. Instead of scaling through programmable infrastructure, diagnostics fragmented into isolated systems—PCR, sequencing, ELISA, culture, flow cytometry—each built around dedicated hardware, analog workflows, and fixed signal architectures. Capability improvements required larger instruments, higher cost, more labor, and increasing operational complexity.
Guanine introduces a software-defined signal architecture for biological measurement.
Within a unified electrochemical signal layer, biological information is generated, encoded, stabilized, and computationally resolved through programmable software rather than modality-specific hardware. Multiplexing, low-abundance detection, time-resolved biology, and distributed deployment become scalable functions of signal density and computation.
This architecture enables:
- Quantitative multi-analyte detection from a single sample
- High-density biological measurement without increasing hardware complexity
- Portable low-cost deployment beyond centralized laboratories
- Real-time biological monitoring and phenotypic analysis
- One platform across molecular, protein, cellular, and dynamic assays
As AI, precision medicine, antimicrobial resistance, distributed testing, and real-time biological intelligence accelerate, existing diagnostic architectures cannot scale to meet demand.
Programmable biological measurement becomes inevitable.
Explore the architectureProgrammable Architectures Changed Information Processing
Modern computing did not scale because transistors became slightly better. It scaled because information processing became layered, programmable, and software-defined. Shared signal architectures separated hardware from capability expansion. Operating systems, instruction sets, algorithms, and software ecosystems unlocked exponential scaling without requiring a new machine for every application.
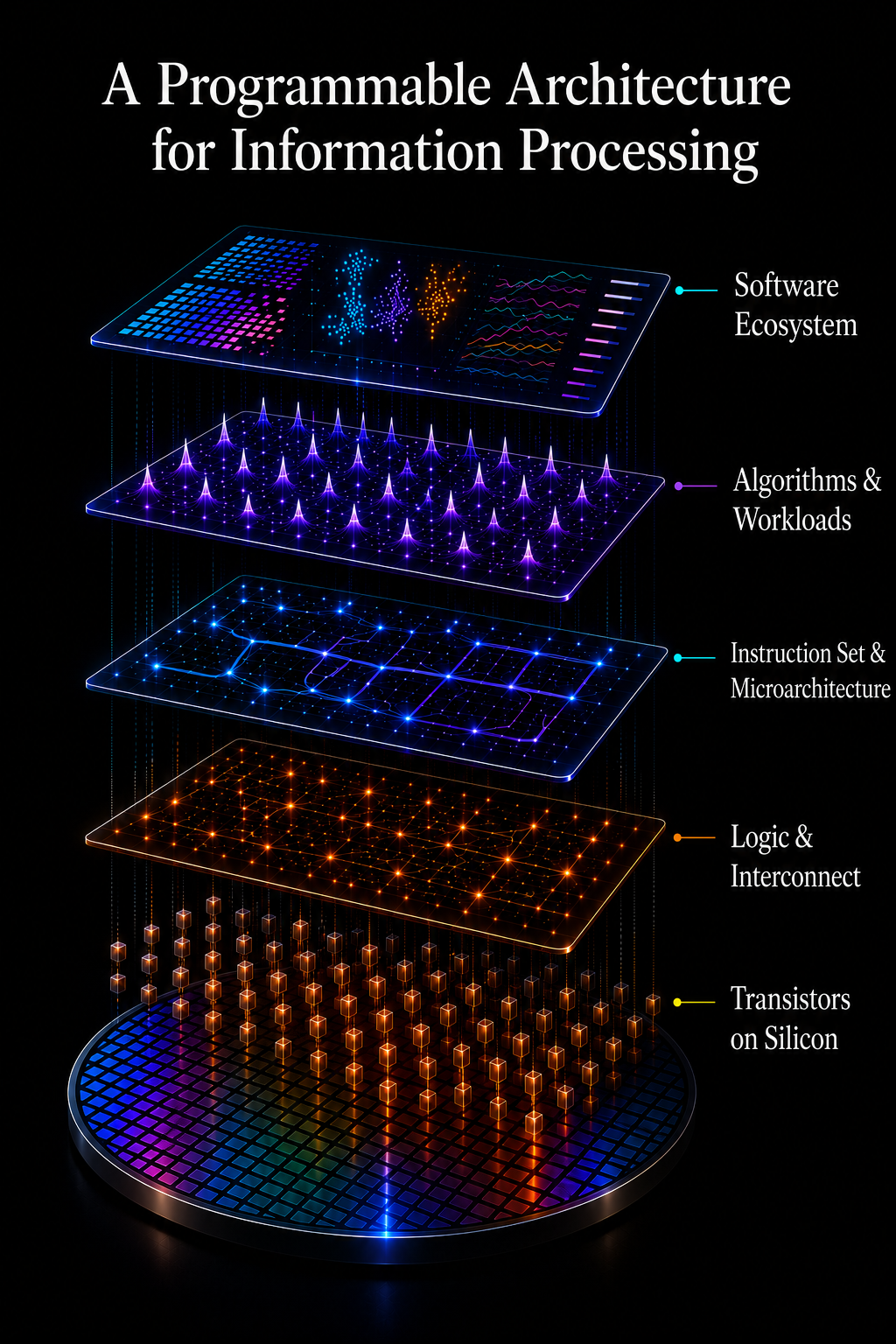

Fragmented Biological Systems Prevent Measurement from Scaling
Biological measurement remains trapped in fragmented analog architectures. Genomics, proteomics, microbiology, and specialized assays each require dedicated instruments, workflows, and signal systems. Capability improvements come through increasing complexity, capital cost, and operational burden rather than through shared programmable infrastructure.
A Programmable Signal Architecture for Biological Measurement
Guanine replaces modality-specific systems with a programmable electrochemical signal stack. Signal generation, encoding, stabilization, and computational interpretation operate as layered software-defined functions within a unified architecture. Biological information becomes digitally addressable, computationally scalable, and deployable across many assay categories without redesigning the core system.
Explore the architecture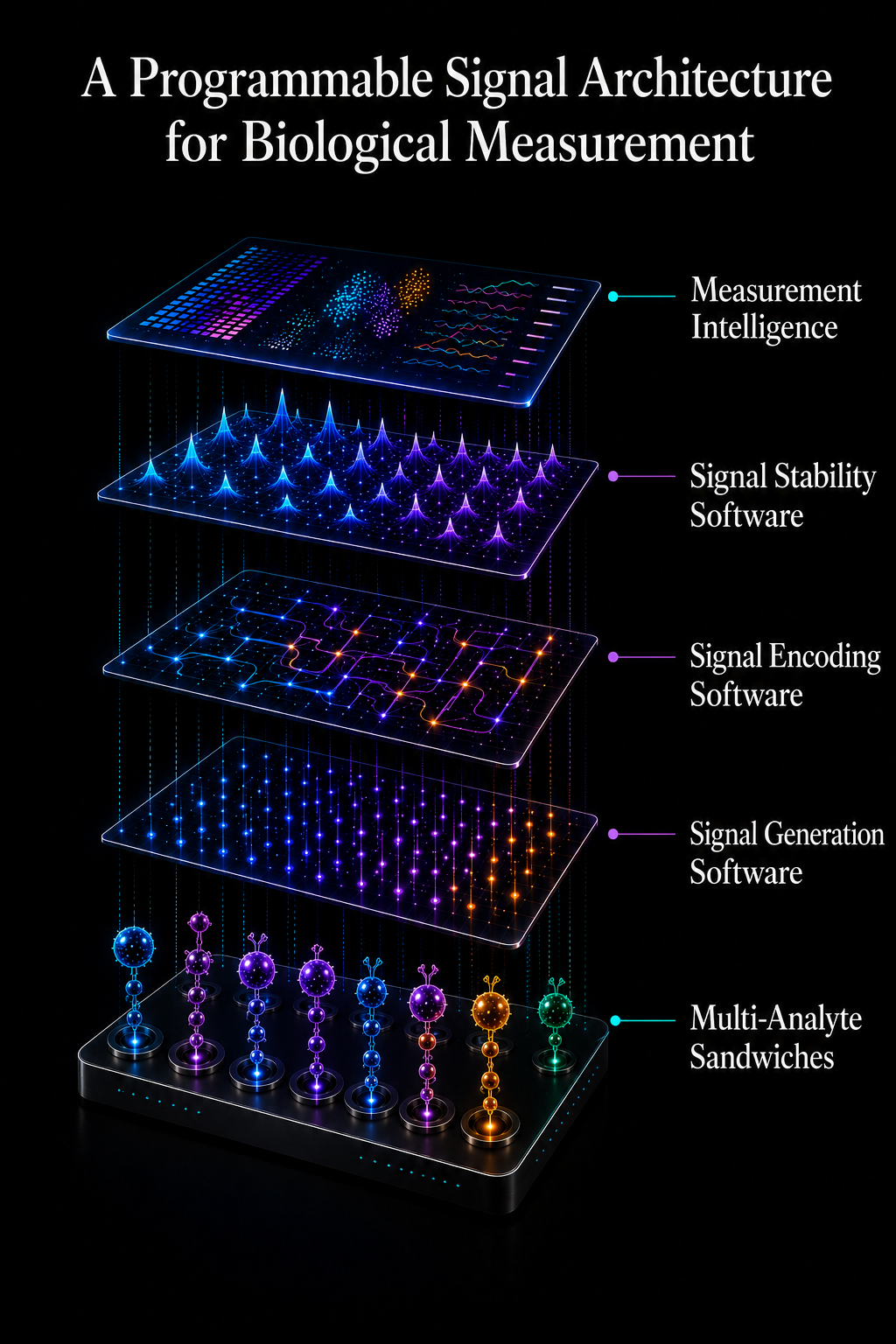
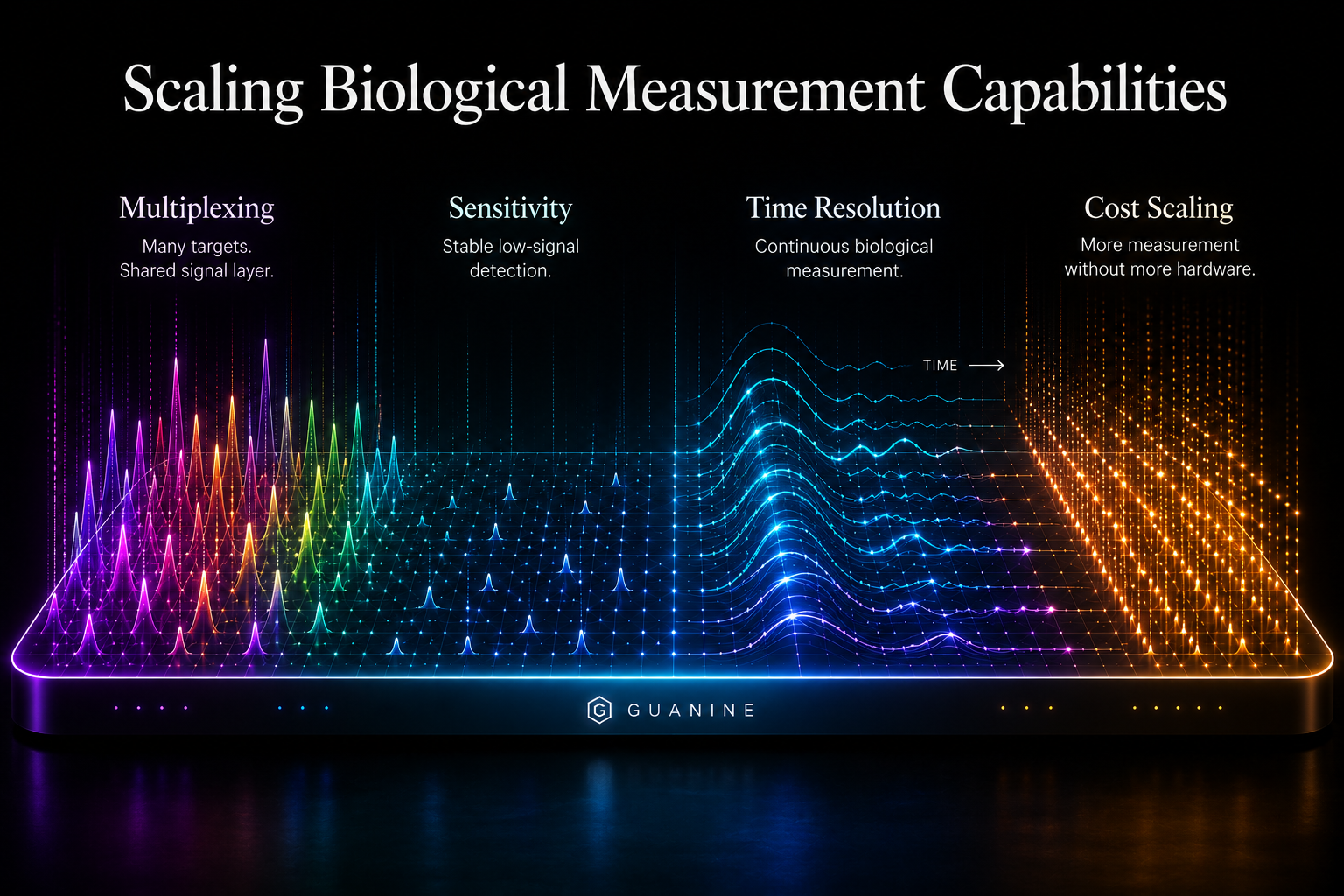
Multiplexing, Sensitivity, Time Resolution, and Cost Scaling
Legacy systems force tradeoffs between multiplexing, sensitivity, time resolution, and deployment cost. Guanine’s architecture enables all four simultaneously within a shared signal system. High-density biological measurement no longer requires larger instruments, longer workflows, or centralized laboratories.
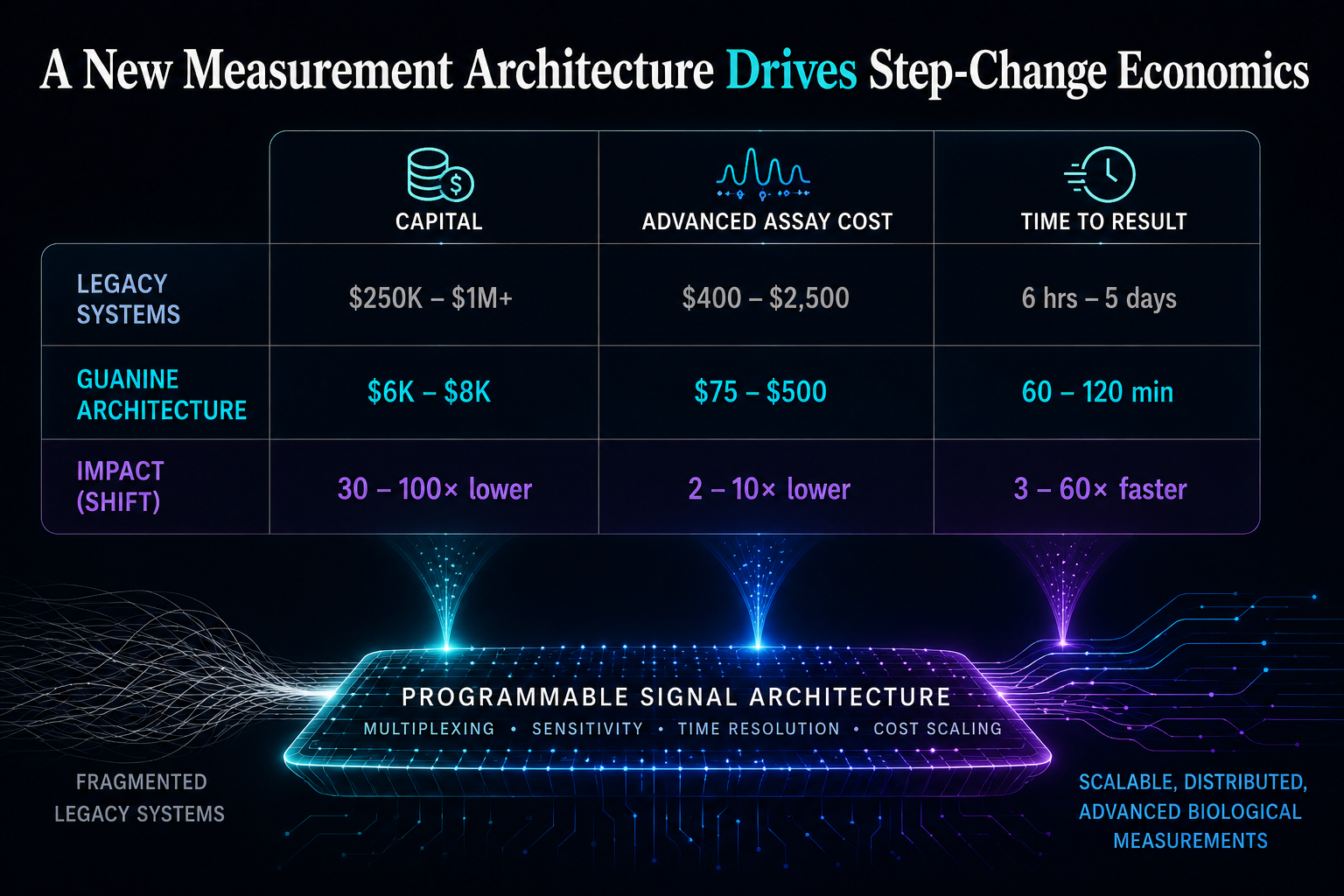
A New Measurement Architecture Drives Step-Change Economics
Advanced biological measurement historically required large centralized instruments costing hundreds of thousands to millions of dollars. Guanine reduces capital requirements while enabling advanced multiplexed assays at dramatically lower operational cost and faster turnaround. Measurement capability scales through software and signal density rather than hardware complexity.

One Architecture Across Many Biological Markets
The same programmable signal architecture supports low-abundance proteins, genetic targets, multi-omic panels, rare pathogens, time-resolved biology, and distributed testing. OEM partners can deploy advanced assay categories on compact low-cost readers without building entirely new instrument ecosystems.
Explore the OEM platformCulture-Free Phenotyping Within the Clinical Decision Window
Sepsis therapy decisions must occur within 1–2 hours, while conventional culture and susceptibility workflows require days. Guanine enables actionable pathogen identification and phenotypic susceptibility within the first clinical decision window using a programmable multiplexed biological measurement system designed for rapid distributed deployment.
See the sepsis workflow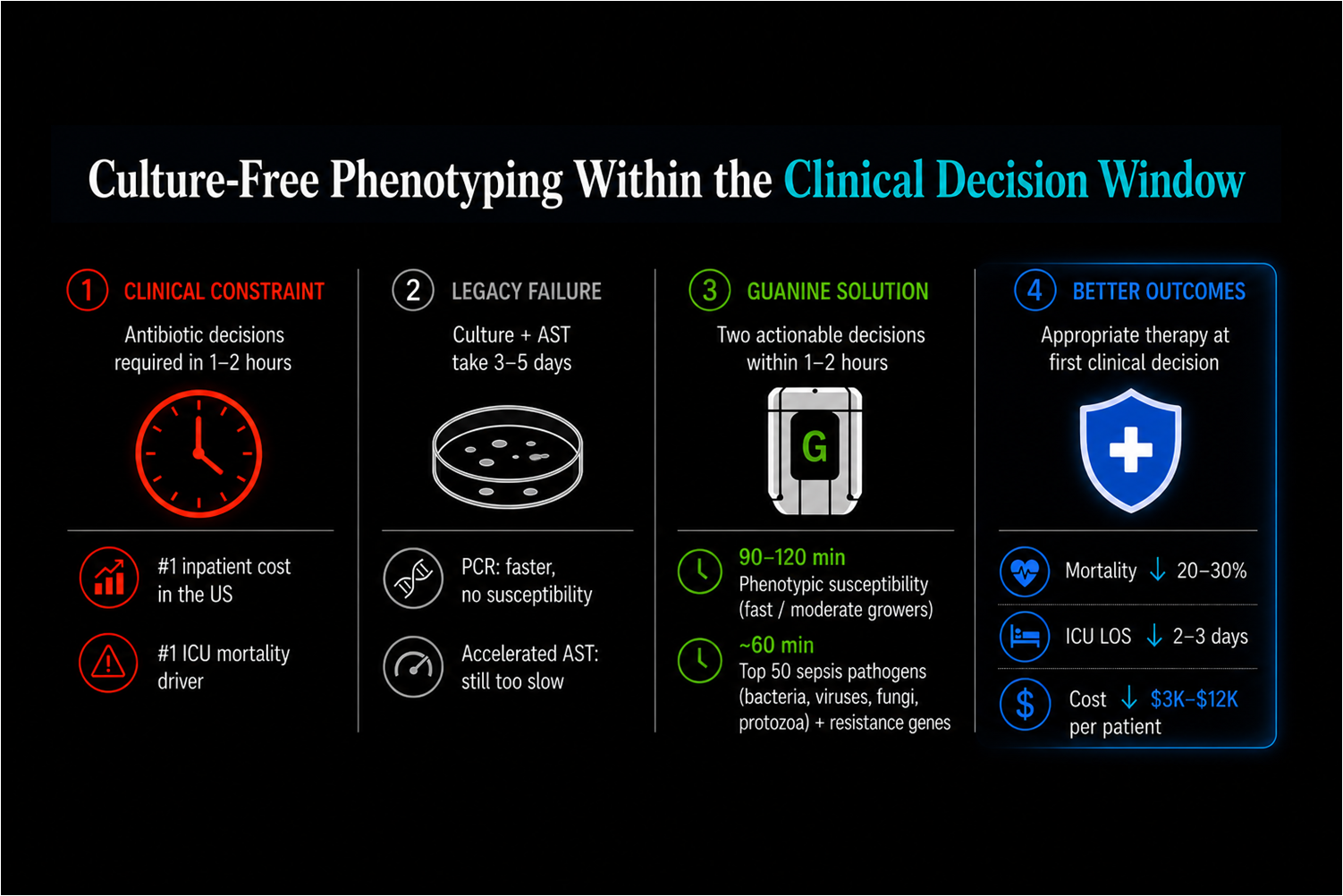
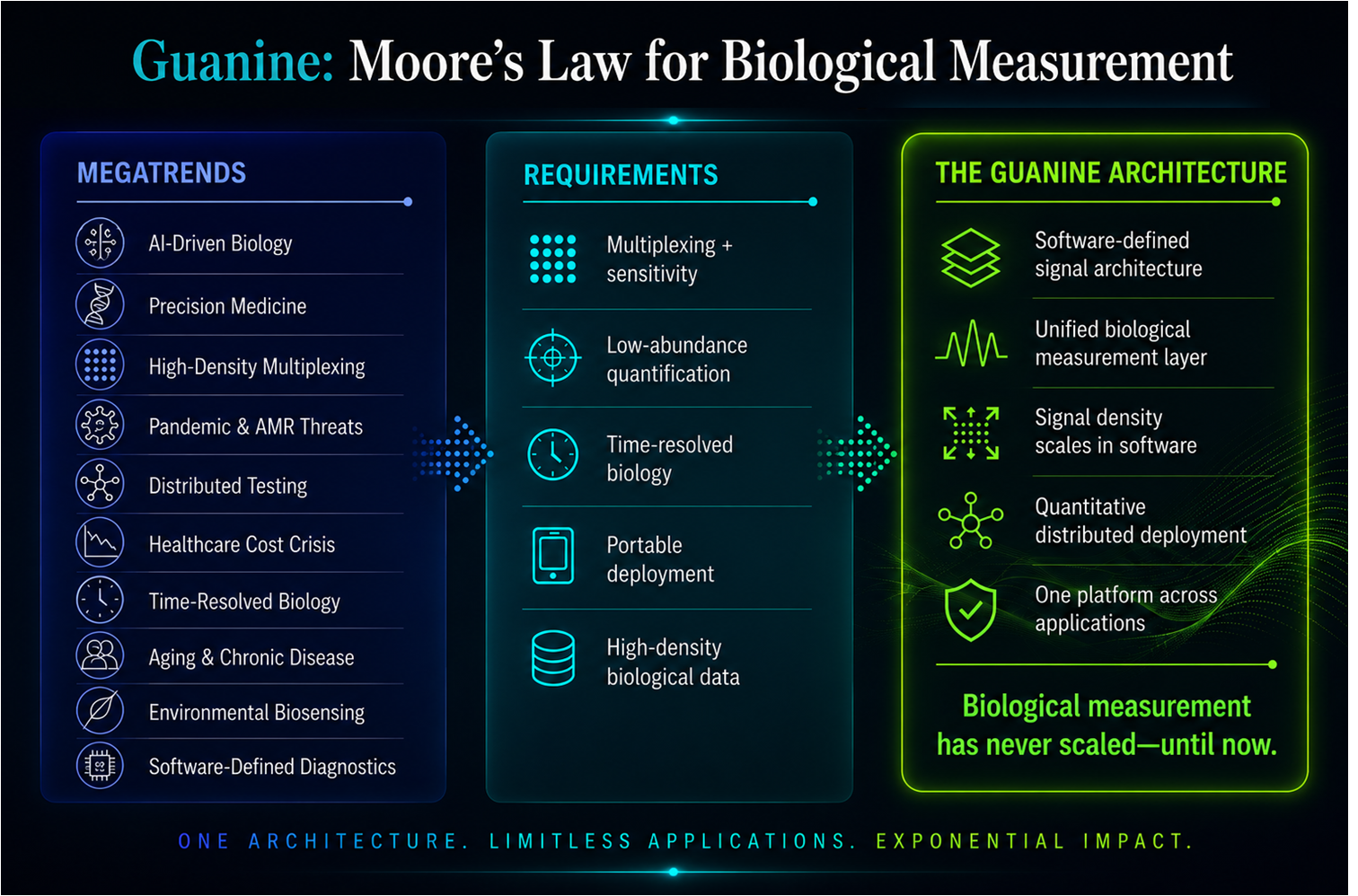
Megatrends Are Driving Demand for Scalable Biological Measurement
AI-driven biology, precision medicine, antimicrobial resistance, distributed diagnostics, environmental biosensing, and real-time biological monitoring are converging into a single requirement: scalable biological measurement. Existing architectures cannot support the required combination of density, sensitivity, deployment, and economics. A programmable signal architecture enables biology to scale like information systems before it.
Contact us for more information
