⭐ A Scaling Law for Biological Measurement
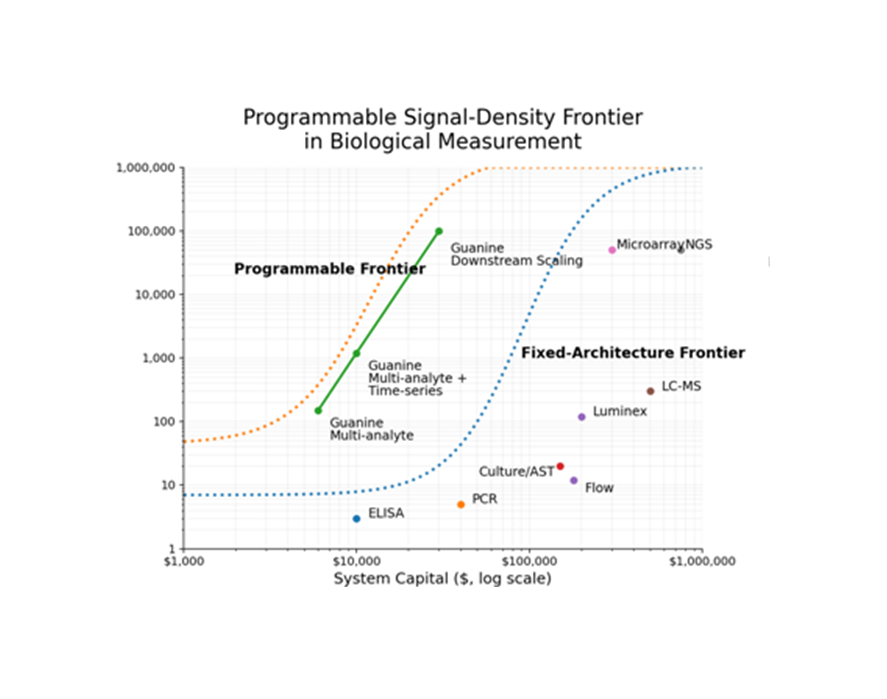
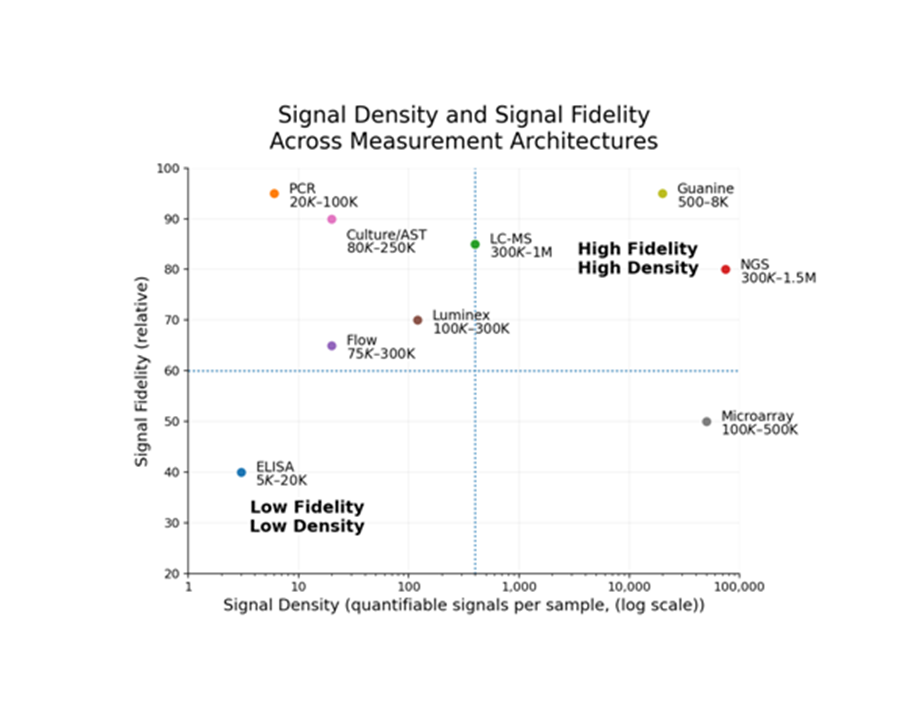
Biological measurement can scale across dimensions
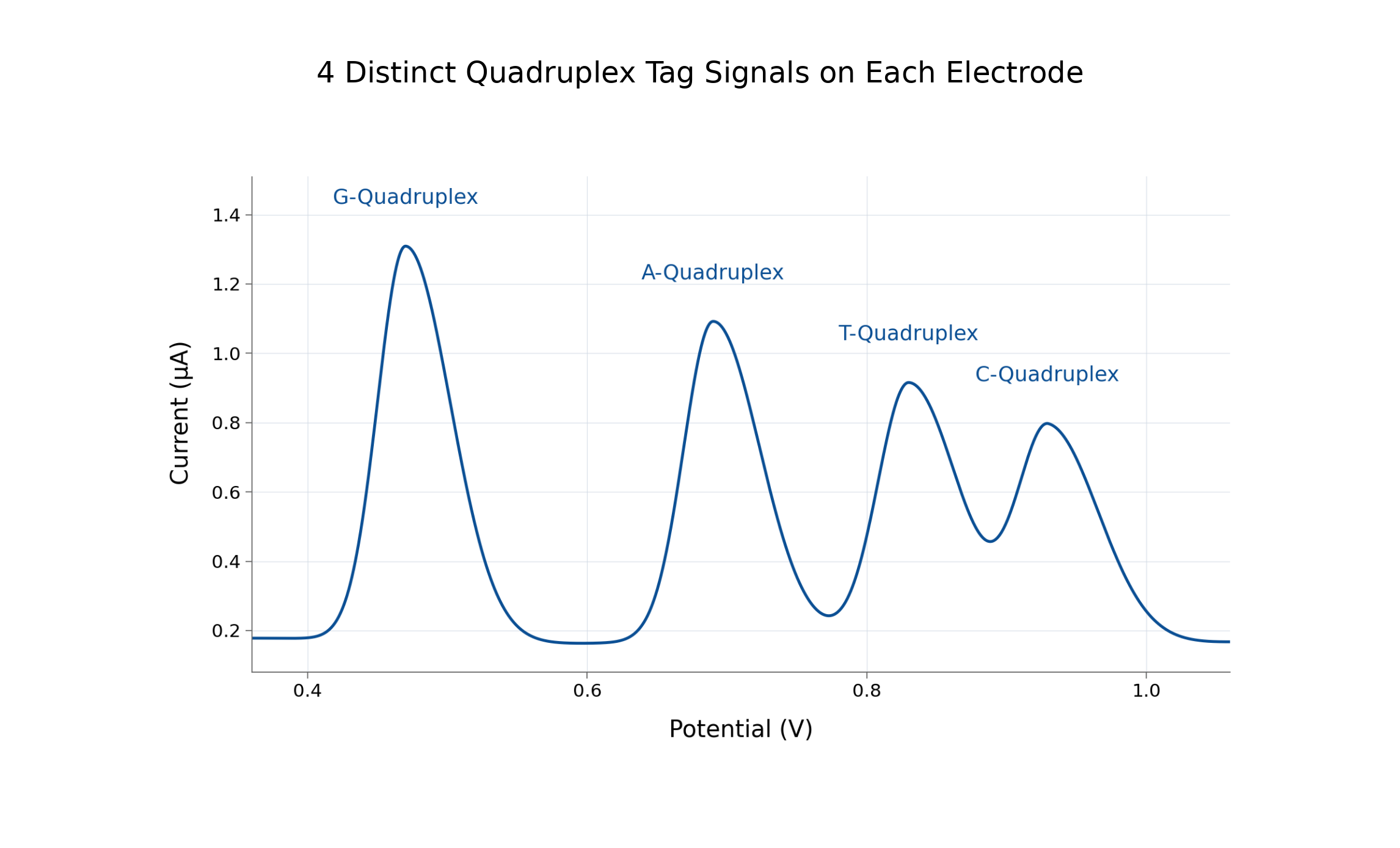
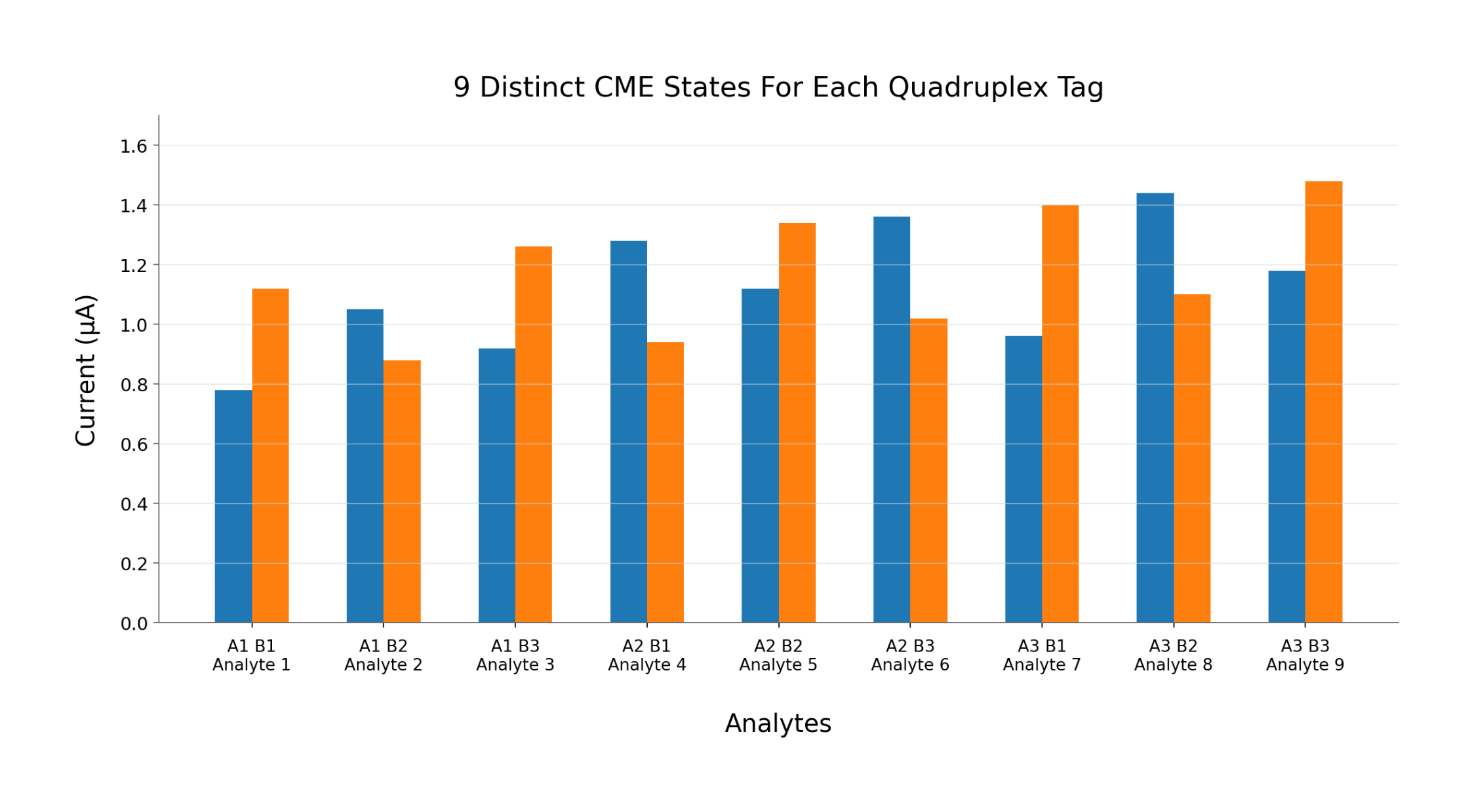
Guanine introduces signal density as the primary scaling mechanism. Each analyte is converted into a dense, encoded electrical signal—measured and resolved through software rather than hardware expansion.
Guanine is built around a different premise:
biological measurement can scale when signal becomes the unit of engineering.
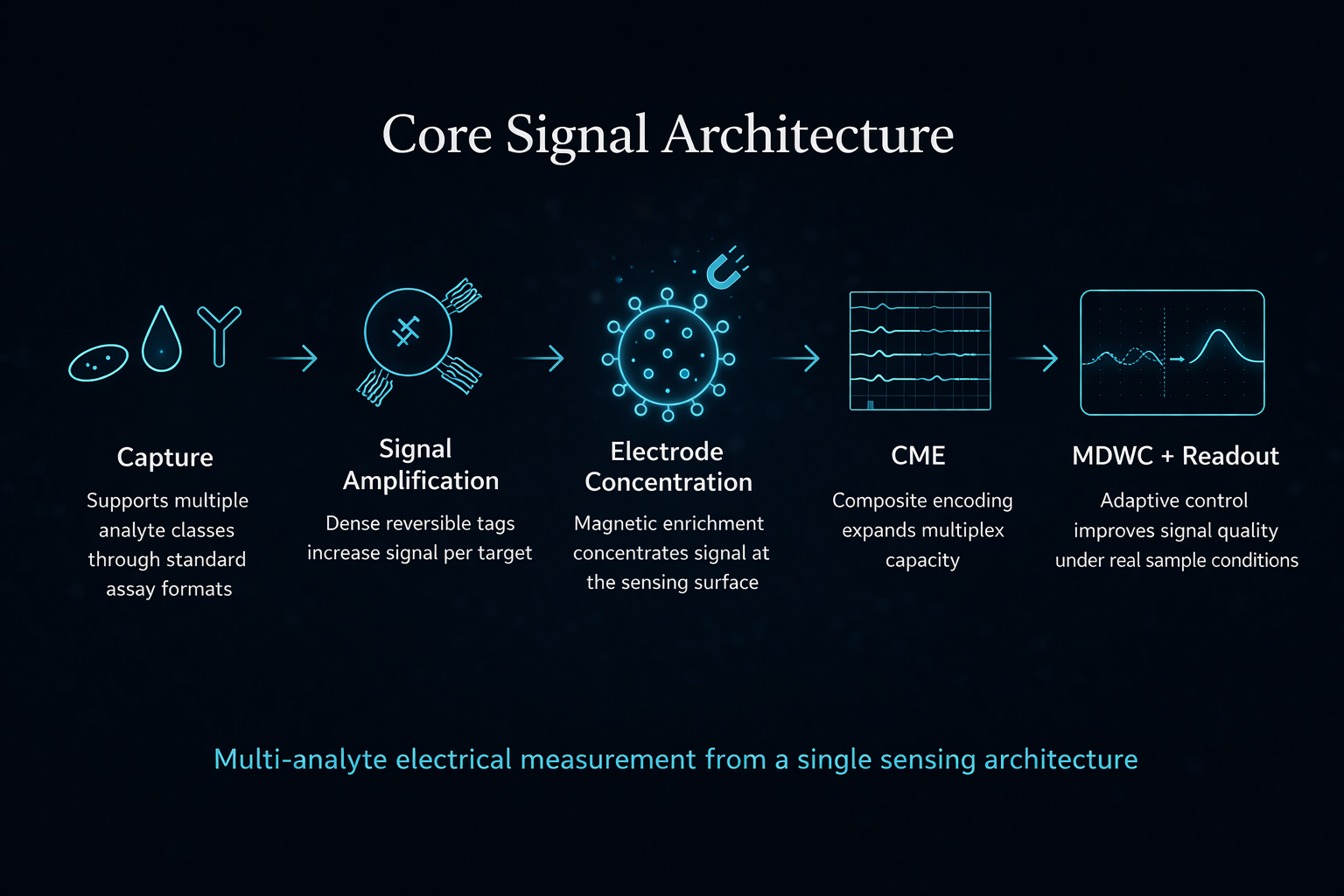
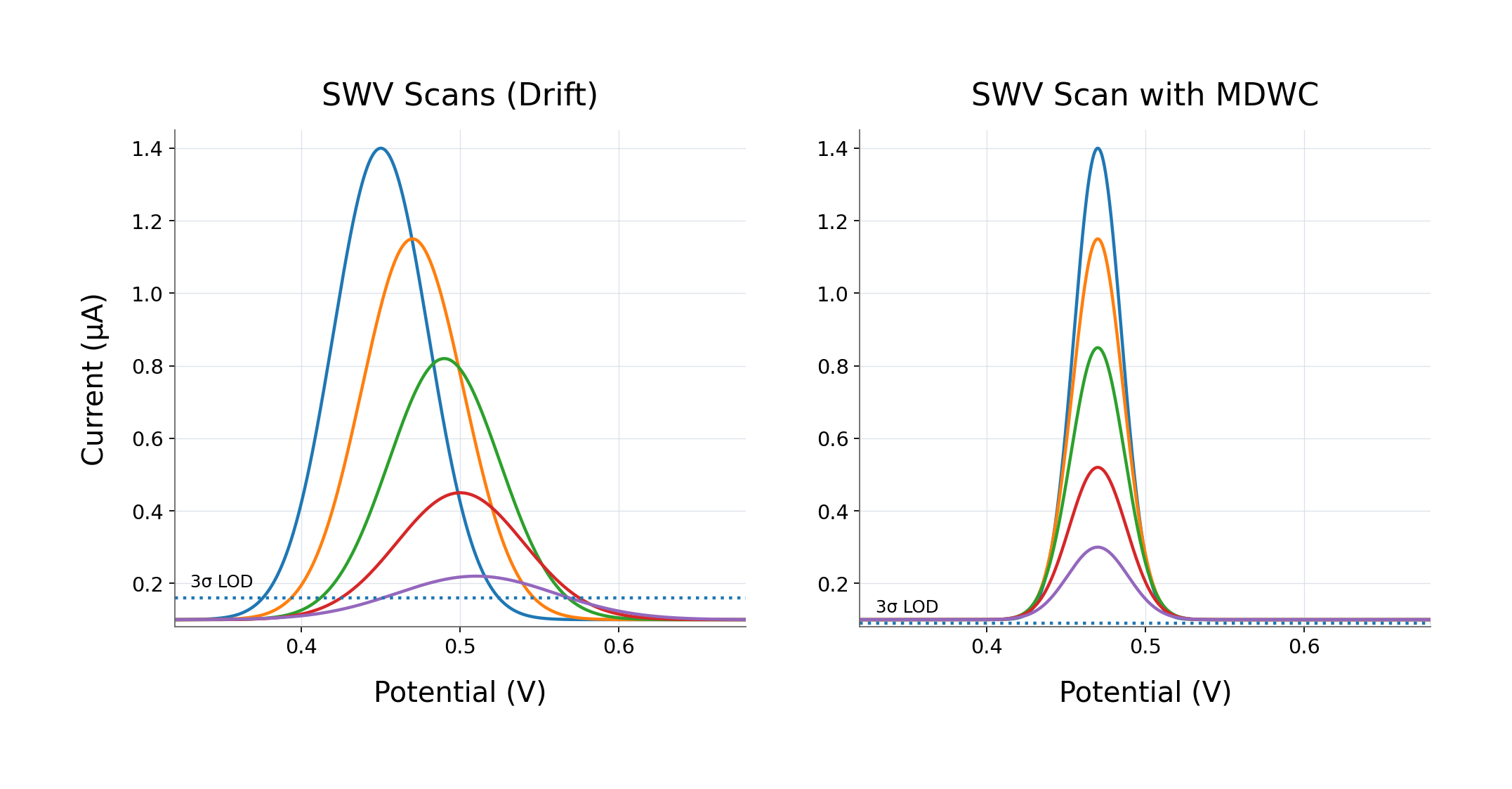
By combining a multi-analyte electrochemical signal primitive, a combinatorial address space for multiplexing, a control layer that stabilizes signal as density increases, and integrated cartridge workflows,
Guanine shifts measurement from chemistry-bound workflows to signal-defined infrastructure.
Legacy systems scale by adding hardware. Guanine scales by increasing signal density.
• Capability increases without adding system complexity.
• Multiplexing expands without new instrumentation.
• Signal strength scales with encoding, not enzymatic amplification.
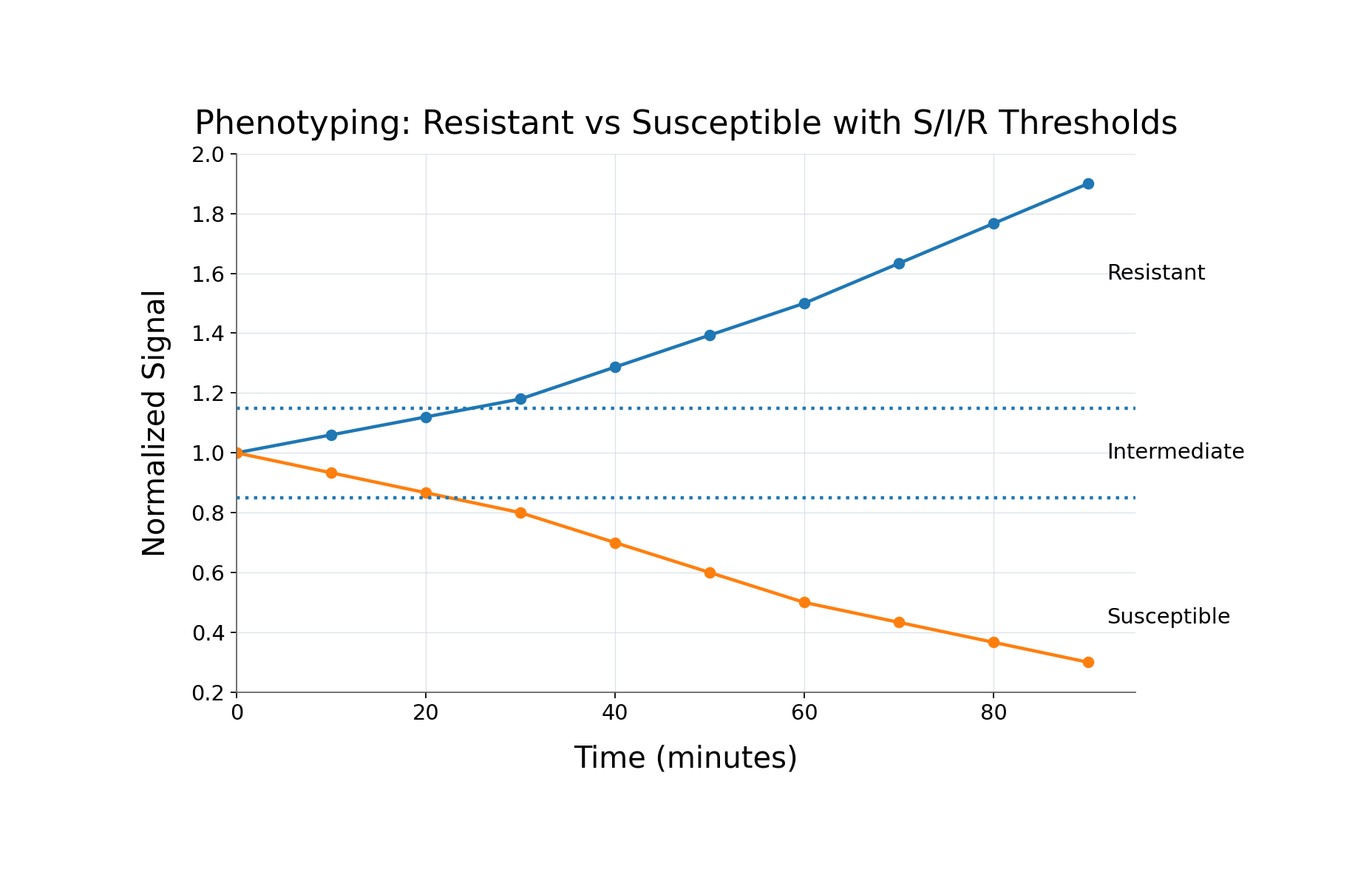
This enables combinations that existing systems cannot achieve: high multiplex with low cost, high sensitivity with rapid turnaround, multi-analyte measurement from a single sample, and functional time-series measurement within a single test.
Not an incremental improvement—a transition from fixed diagnostic systems to programmable measurement infrastructure.
Moore’s Law was not just smaller components. It was a system architecture that allowed capability to compound.